
Once every five days. That’s how often our researchers have published papers in peer-reviewed scientific journals, on average, since the ARC Centre of Excellence in Plant Energy Biology (PEB) was founded in 2005.
The world-leading Australian plant research Centre recently hit a milestone 1000 publications, representing 1000 significant contributions to plant and related sciences
To celebrate, we’re recounting some of the biggest, weirdest, most impactful and most surprising discoveries of the last 15 years.
We’ll be sharing some of our discoveries throughout November,
showcasing research into salinity, epigenetics, stress, drought, plant proteins
and more. Keep your eye on this page and our social media channels!
PEB Director Professor Harvey Millar said it was incredible to mark such a significant achievement.
“A thousand papers is a remarkable number of quality publications for a research Centre in just 15 years.”
“We’re proud to have been able to make a huge contribution to the field of plant science,” he said.
The 1000 PEB publications have, collectively, clocked over 80,000 citations by other researchers from around the world, having far reaching impact in the research and general community alike.
“From top publications in leading journals such as Cell and Science to quirky findings that captured people’s imagination, each paper marks a unique and significant step in our quest to understand more about the world around us.”
PEB is focused on understanding the way plants capture, convert and use energy in response to environmental change, with a view towards improved plant energy efficiency.
Professor Millar said the Centre’s research has big impacts in food security, agriculture and climate adaptation.
“We largely use taxpayers’ money as we explore the world of plants around us,” he said.
“It is essential we honour the trust society has placed in us and use our discoveries to make a difference in the immediate society we live in, and the global society we depend on.”
PEB has seen hundreds of plant scientists working together across Australia over the last 15 years, with research nodes at the University of Western Australia, the University of Adelaide, the Australian National University, La Trobe University and, historically, Flinders University and the University of Sydney.
A
full list of PEB’s publications can be viewed here. Follow #1000discoveries across social media on our Twitter, Facebook and Instagram channels.
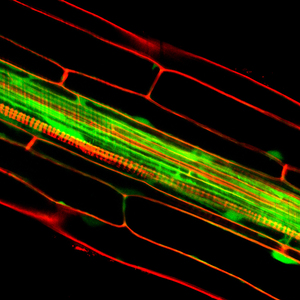

Discovery #357: How plants react when there isn’t enough phosphate
2013 in THE PLANT CELL.
Discovering how plants respond to phosphate deficiency was the first step in training them to use less fertiliser.
In this paper, PEB researchers studied how rice plants respond when they don’t have enough phosphate, an essential nutrient for life and a major component of chemical fertilisers. The team profiled the response of all rice genes to phosphorus, and uncovered a large number of potential regulators of phosphate in plants.
These new leads represented a major step forward in understanding how plants respond to nutrient deprivation, and opened up a whole new area of processes we hadn’t been aware of before.
Following this research, PEB scientists have successfully re-engineered the DNA of plants to make them hardier and less dependant on phosphate. They’ve been able to manipulate gene networks to ‘trick’ plants into thinking phosphate is scarce so that they use the phosphate available to them more efficiently.
If translated to crop plants, the research could save Aussie farmers $300 million a year and significantly reduce run-off phosphorous in waterways. The work will also help to boost food security amid diminishing global phosphate supplies.
View the full paper.

Discovery #39: The global go-to for the location of proteins
2007 in NUCLEIC ACID RESEARCH.
A database for model plant Arabidopsis has become a research tool of choice by plant scientists around the world.
PEB scientists pooled data on the location of proteins in the cells of Arabidopsis thaliana, a small, rapid-growing cress that is used as a model plant by researchers all over the world. They shared the results in an online database that has since become a widely-used research tool for the plant science community.
Known as SUBA, the database pools experimental data and predictions from hundreds of studies to create a powerful research tool. It allows anyone to access the sub-cellular location of proteins in Arabidopsis, as well as the interactions between different proteins. SUBA has undergone several iterations over the years, with the current release being SUBA4.
PEB later created a similar database for economically-important crop plants, called CropPAL. It pools ten years of research by more than 300 institutions into a significant resource for crop improvement and global food security. The database is accelerating solutions that address challenges such as drought, salinity and nitrogen starvation in major global crops.
View the full paper.
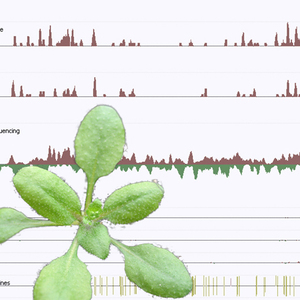
Discovery #83: The world’s first integrated high-res epigenome map
2008 in CELL.
This landmark study showed the world how to map ‘on-off switches’ for genes.
PEB helped to pioneer the mapping of the epigenome—the molecular code that can control which regions of our genes are expressed at any given time. Controlling when and where genes are turned on is a major factor in cell identity, for example what makes a skin cell different to a brain cell, even though they have the same underlying DNA.
Just as the Human Genome Project was the first to catalogue all the genes in the human body, this study achieved several firsts in the high-resolution cataloging of the epigenomes of living organisms – and did so in a plant! The research team used the model plant Arabidopsis thaliana, whose genome is 25 times smaller than humans.
The paper described two new genomic methods. The first enabled the high-resolution mapping of the Arabidopsis epigenome. It works by precisely determining whether each C base in the genome is methylated or not, and then layering the resulting epigenomic map onto the exact genome it regulates. The second allowed mapping of expressed genes by a sequencing method that is now known as RNA-seq and used extensively by researchers worldwide.
The technique was later used to generate the first maps of the human DNA methylome - a critical layer of the epigenome - in research rated by TIME magazine as the second most important scientific discovery of 2009. The work has helped us understand human diseases, such as cancer, as well as how the epigenome changes because of our age, diet and environment.
View the full paper.

Discovery #544: Plants everywhere respond to heat in the same way
2016 in PROCEEDINGS OF THE NATIONAL ACADEMY OF SCIENCES.
From Western Australia to Alaska to Costa Rica; plants react the same when it’s hot.
A study of vegetation at 18 remote sites around the world found striking similarities in how different plants respond to increasing temperatures. The research looked at the rate of plant respiration—a process in which plants take in oxygen and release carbon dioxide.
Existing climate models worked on the basis that respiration doubled for each 10°C rise in temperature. Instead, this research study found respiration is more temperature sensitive than previously assumed, and becomes less sensitive as the temperature rises. This pattern plays out across the globe.
The team studied plants growing in seven different habitats, from the arid woodlands of Western Australia to the deciduous forests of New York, the arctic tundra in Alaska, the boreal forests of Sweden and the tropical forests of Costa Rica and Peru.
Despite the diversity of plant types, there were striking similarities in how different plants alter their respiration rate in response to increasing temperature. The research was the most comprehensive study of plant respiration responses to temperature ever conducted.
View the full paper.

Discovery #633: Teaching plants to use their energy more wisely
2017 in THE PLANT CELL.
Knowing exactly how much energy plants need to maintain their proteins could lead to more energy-efficient crops.
In this study, PEB researchers used a new technique to calculate the amount of energy plants need to maintain their proteins, one of the most energy-intensive processes for plants.
To grow and maintain themselves, plants must constantly create new proteins and break down existing ones. The process is called ‘protein turnover’, and it uses much of a plant’s energy.
The team were able to calculate exactly how much energy is needed for more than 1000 different proteins found in leaves. They measured how long each protein lives, and how much energy plants need to spend to keep that protein around and functional.
Some proteins lasted just a few hours, while others stuck around for months. Identifying features of long-lasting proteins could help us engineer more robust, less energy-expensive proteins. If we can teach plants to use their energy wisely in this way, we can create more efficient and productive crops.
View the full paper.

Discovery #538: Plants usually forget stressful events
2016 in SCIENCE ADVANCES.
Plants, it seems, prefer not to hold grudges.
Studies have shown that plants are able to ‘remember’ stressful events such as droughts, helping them to adapt and survive similar conditions in the future. Because the process involves DNA, there is also evidence that plants can pass long-term ‘memories’ to their offspring.
But there are relatively few examples of these memories. And when PEB researchers scoured the literature this review, they found that plants remembering stressful events tended to be the exception rather than the rule. In short: plants are generally good at forgetting.
The team hypothesised that whether a plant forms a memory likely depends on what happens after a stressful experience. During this ‘recovery phase’, the plant can either consolidate its stress response and remain genetically primed, or reset to its prior state.
The researchers tested their ideas in a subsequent study looking at how plants recovered from stressful weather in the lab. They put plants under light stress for an hour and then allowed them to recover for an hour, supporting the work with mathematical modelling.
The team found plants are able to recover phenomenally well from some environmental stresses, quickly resetting to the pre-stress state. The findings could help us better understand how plants and crops will cope and recover from variable weather.
View the full paper.

Discovery #243: Salt-tolerant wheat
2012 in NATURE BIOTECHNOLOGY.
Introducing an ancient gene into commercial wheat crops boosted harvests by 25 per cent.
This research was the first in the world to fully describe the improvement in salt tolerance of an agricultural crop — from understanding the function of salt-tolerant genes in the lab to demonstrating increased grain yields in the field.
PEB researchers bred salt tolerance into a variety of durum wheat, improving grain yield by 25 per cent on salty soils. The team first looked to wild relatives of modern-day wheat, discovering a new salt-tolerance gene in an ancestral cousin called Triticum monococcum.
They then used 'non-GM' crop breeding techniques to introduce the ancient gene into commercial durum wheat. They also conducted field trials across Australia, including at a commercial farm in New South Wales.
Salinity affects more than 20 per cent of the world's cultivated land, with predictions that this will double by 2050. This is a particular issue in Australia’s prime wheat-growing areas. It’s a problem because if sodium starts to build up in the leaves of plants, it affects important processes such as photosynthesis. The salt-tolerance gene in the study (known as TmHKT1;5-A) works by producing a protein that excludes sodium from the leaves.
View the full paper.
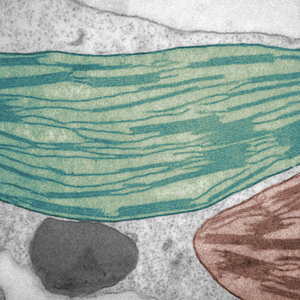
Discovery #229: A new form of plant communication
2011 in THE PLANT CELL.
How plant cells use chemical signals to help beat drought.
PEB researchers discovered a new form of cellular communication used by plants to respond to drought. The team discovered evidence of a process called ‘retrograde signalling'—where chemical signals move between different compartments in plant cells. These signals activate a defence mechanism that could help plants cope with drought.
The researchers examined the model plant Arabidopsis thaliana, a relative of canola. They found that in drought conditions, the chloroplasts—the part of a plant cell responsible for photosynthesis— communicated with the nucleus—the control centre of the cell. This changed the activity of thousands of genes to cope with drought stress.
The study found evidence that a chemical called PAP acts as a signal within the cell. This type of ‘cross-talk’ between cellular compartments also occurs in animals, using other chemicals. In Arabidopsis, the process was helped when a protein called SAL1 was removed from the plant. The research has the potential to boost drought tolerance in commercial crops through selective breeding.
View the full paper.

Discovery #460: Stressed out plants send animal-like signals
2015 in NATURE COMMUNICATIONS.
An important signalling molecule in the human brain — GABA — may be just as essential in plants.
PEB scientists discovered that plants may use a chemical normally associated with animals, when they encounter stressors such as drought, salinity, viruses, acidic soils or extreme temperatures.
The study found that plants respond to stress caused by their environment with a similar combination of chemical and electrical responses as animals do, despite not having a nervous system. But this response happens through systems specific to plants.
We’ve known for a long time that stressed plants produce gamma-aminobutyric acid (GABA), a chemical best known as a neurotransmitter in animals, but there was little evidence that plants use GABA as a signal. This study discovered that plants respond to GABA in a similar way to animals - via electrical signals - and that this controls plant growth.
Researchers are now investigating whether the discovery can be used to breed more stress-resilient crops. The finding could also explain why particular plant-derived drugs work in humans, and opens the door to future plant GABA-signalling research that benefits human medicine.
View the full paper.

Discovery #750: Cracking the genome of bread wheat
2018 in SCIENCE.
An international consortium of more than 200 scientists achieved what was once considered impossible—sequencing the genome of bread wheat.
PEB scientists were part of a global effort to publish the genome of bread wheat, the world’s most widely cultivated crop. Led by the by the International Wheat Genome Sequencing Consortium, the research involved more than 200 scientists from 73 research institutions around the world.
After 13 years of work, the consortium produced the most comprehensive, annotated genome sequence to date for wheat. PEB researchers worked on the PPR gene family, which is one of the largest gene families in land plants and is important for plant growth.
Sequencing the bread wheat genome had long been considered impossible because of its enormous size—five times larger than the human genome. The genome is also extremely complex, with three sub-genomes and more than 85 per cent composed of repeated elements.
Wheat is the staple food of more than a third of the global human population, accounting for almost 20 per cent of the total calories and protein consumed worldwide—more than any other single food source. To meet the demands of the world’s projected 9.6 billion people in 2050, wheat productivity needs to increase by 1.6 per cent each year.
The high-quality, annotated genome sequence gives breeders new tools to develop wheat varieties better adapted to climate change, with higher yields, enhanced nutritional quality and improved sustainability.
View the full paper.
